 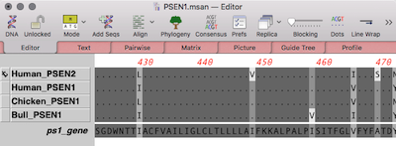
Easy to use for anybody familiar with Windows. If you can use a Mac, you can analyze your sequences with MacVector! So we wanted MacVector for Windows to be the same. MacVector is designed for, and fully integrates with, the OS X environment. It is a fully native version of MacVector but on Windows. MacVector for Windows is not a Java version of MacVector. It’s not yet ready for release, but MacVector for Windows is very definitely on its way. We have a blog with contributions from a selection of MacVector, Inc staff discussing a variety of subjects from OS X through bioinformatics, to the challenges of running a small multi-national company.It’s been one of the most popular customer requests for a long time. There's a useful Getting Started Guide to help new users get going with MacVector. Ĭheck out our promotions page to see the latest deals on MacVector software Which edition is right for you? Here's a handy functional comparison chart. Click here for details of the Cloning Edition. There is a cost-effective version of MacVector available, targeted at users who want the power of MacVector's graphical annotation, clone construction and primer design tools, but without all of the bells and whistles of the full version. So why not upgrade?Ĭheck out this link for a list of the new features introduced in each version going all of the way back to MacVector 10.0. Earlier versions of MacVector can't do this. MacVector 17 and later are not only fully supported and compatible with Apple's latest OS release, macOS Big Sur, but also support Dark Mode and use the latest Apple signing and notarization procedures so you can be sure you are using an authentic copy. The f ull installer can be found on this page. You can read about the new functionality and check your eligibility. Otherwise, it is essentially identical to MacVector 18.0. MacVector 18.1 is a Universal Binary, capable of running natively on both Intel and Apple Silicon Macintosh computers. The full installer can be found on this page. 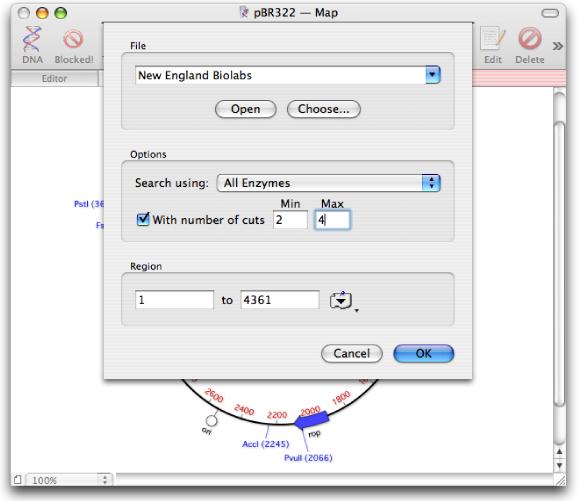
You can read about the new functionality and c heck your eligibility. MacVector 18.2 contains enhancements to the Align to Reference interface and algorithm, improvements to the display of context-sensitive menus and the ability to create Primer Database files from Excel data. Check if you are eligible for the new release and then download the full installer. Note that MacVector Free does require a Macintosh computer running OS X 10.9 or later.Īlong with the usual collection of bug fixes and minor enhancements, MacVector 18.5 has a new heterozygote analysis and base-calling function to find and report mixed residues in Sanger sequencing files. It doesn't have all the advanced bells and whistles of the full release, but it might be just what you are looking for. 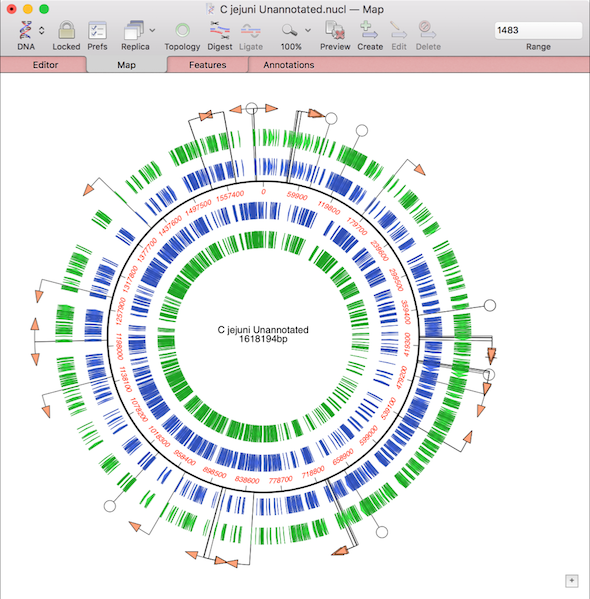
If you have relatively minimal sequence editing and analysis needs, or you just want to view, print and/or convert sequence files sent to you by a colleague, do check out our new MacVector Free product. Assembler must be purchased separately from MacVector. More information on MacVector.Īssembler is an add-on DNA sequence assembly module for MacVector that provides a simple graphical interface to the phred, phrap and cross_match contig assembly algorithms from the University of Washington, the popular Bowtie fast reference alignment program for Next Generation Sequencing projects and the Velvet, SPAdes and Flye de novo assemblers for NGS data. MacVector is widely regarded as the most intuitive, easy to use program available for sequence analysis. MacVector is a comprehensive Macintosh sequence analysis application that provides sequence editing, primer design, internet database searching, protein analysis, sequence confirmation, multiple sequence alignment, phylogenetic reconstruction, coding region analysis, agarose gel simulation and a variety of other functions. develops powerful but easy to use Macintosh applications for Molecular Biologists to simplify and speed up the analysis, manipulation, assembly and documentation of DNA and protein sequences. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
